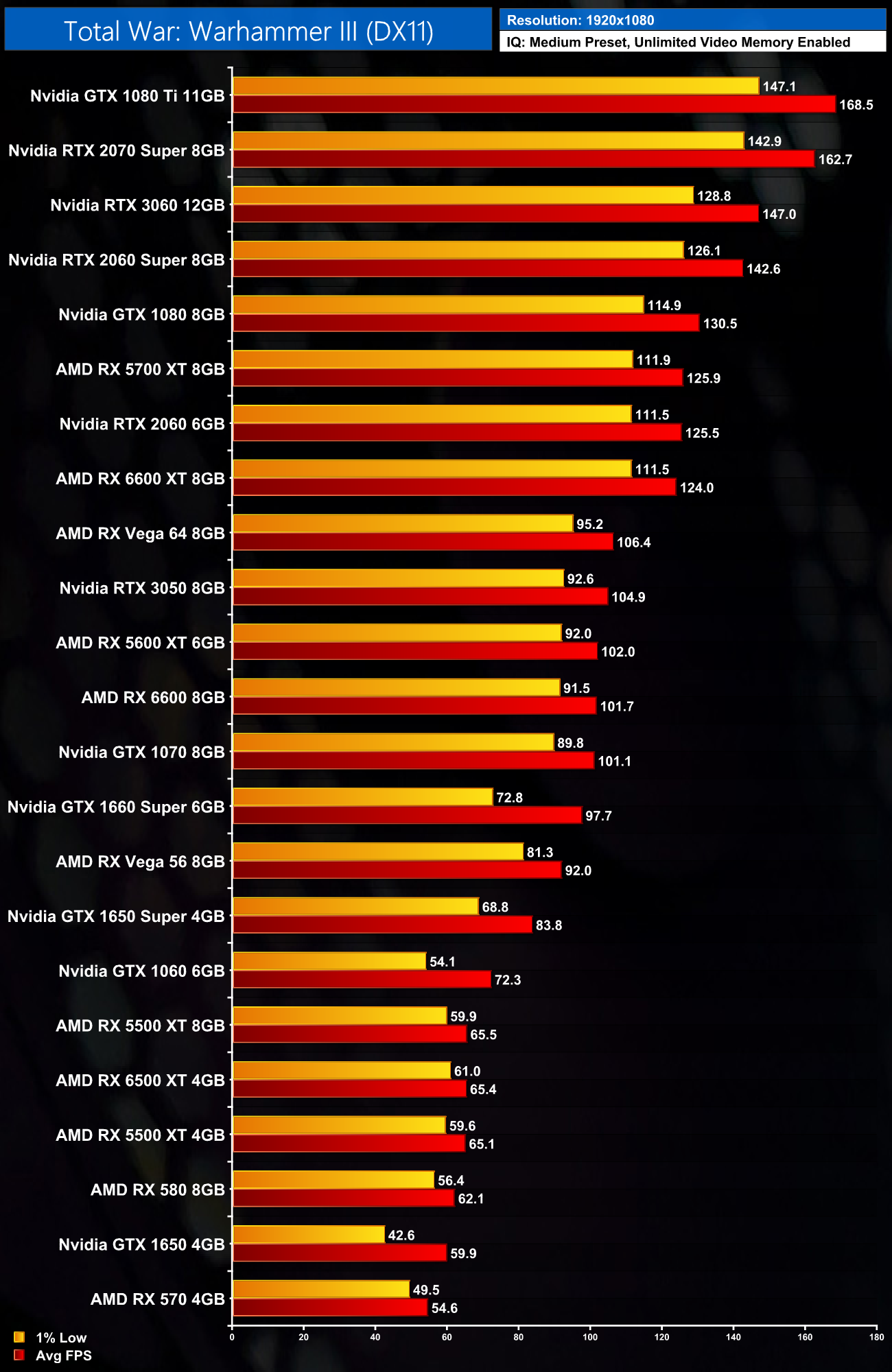
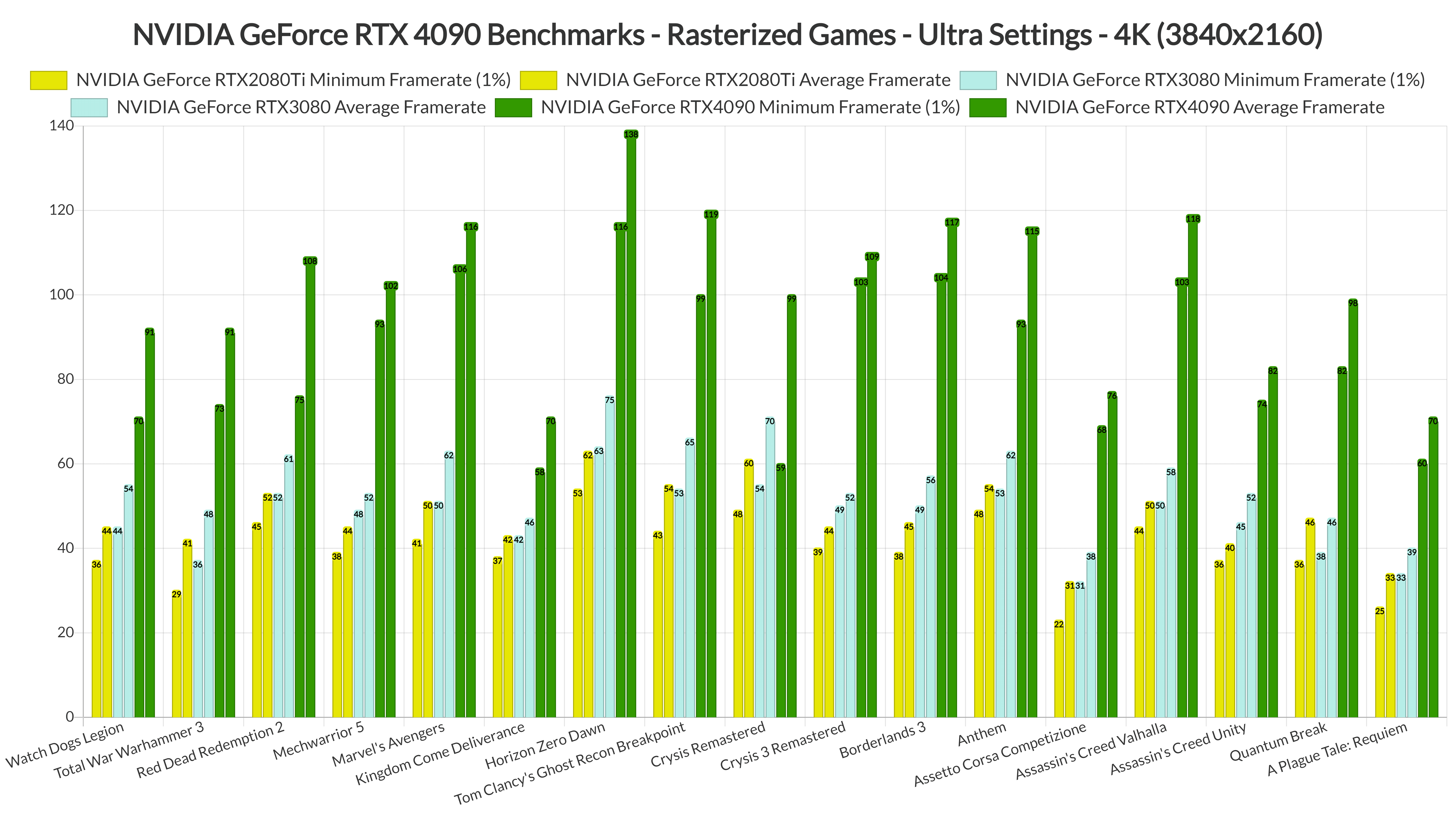
The pmemd.cuda GPU Implementation
Por um escritor misterioso
Descrição

GUI and output for statistical comparison of dynamic motion

AMBER14 & GPUs

Statistical machine learning for comparative protein dynamics with the DROIDS/maxDemon software pipeline - ScienceDirect

Full article: Solvated and generalised Born calculations differences using GPU CUDA and multi-CPU simulations of an antifreeze protein with AMBER
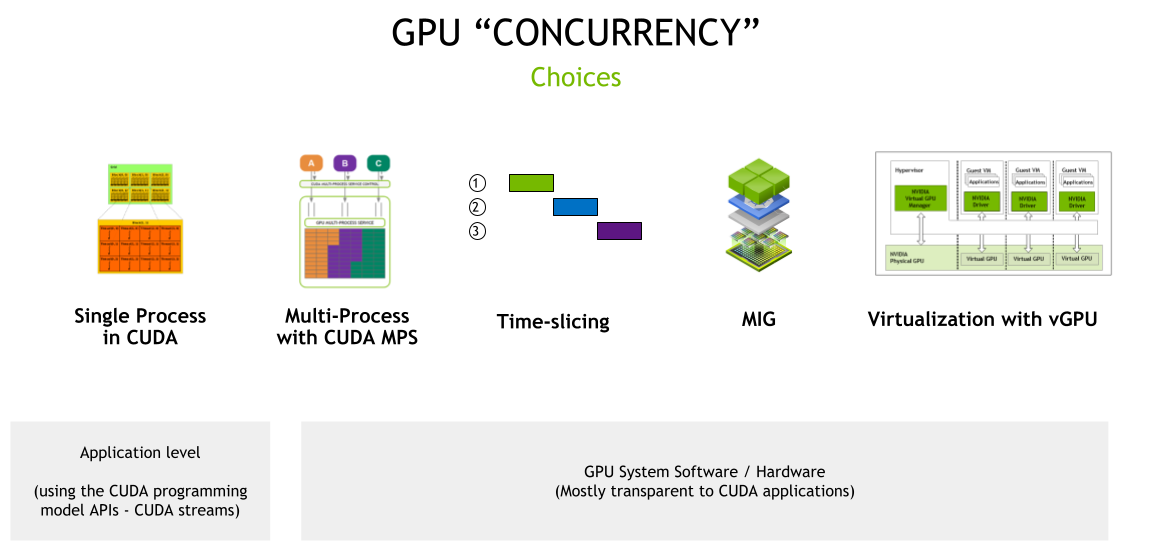
Improving GPU Utilization in Kubernetes
The pmemd.cuda GPU Implementation
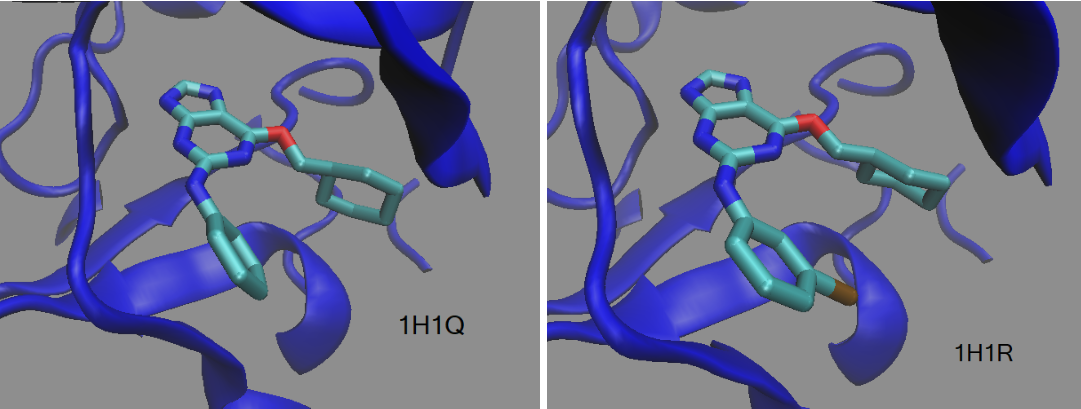
TI Calculation with DDBoost

GPU-Accelerated All-Atom Particle-Mesh Ewald Continuous Constant pH Molecular Dynamics in Amber

Automated docking refinement and virtual compound screening with absolute binding free energy calculations
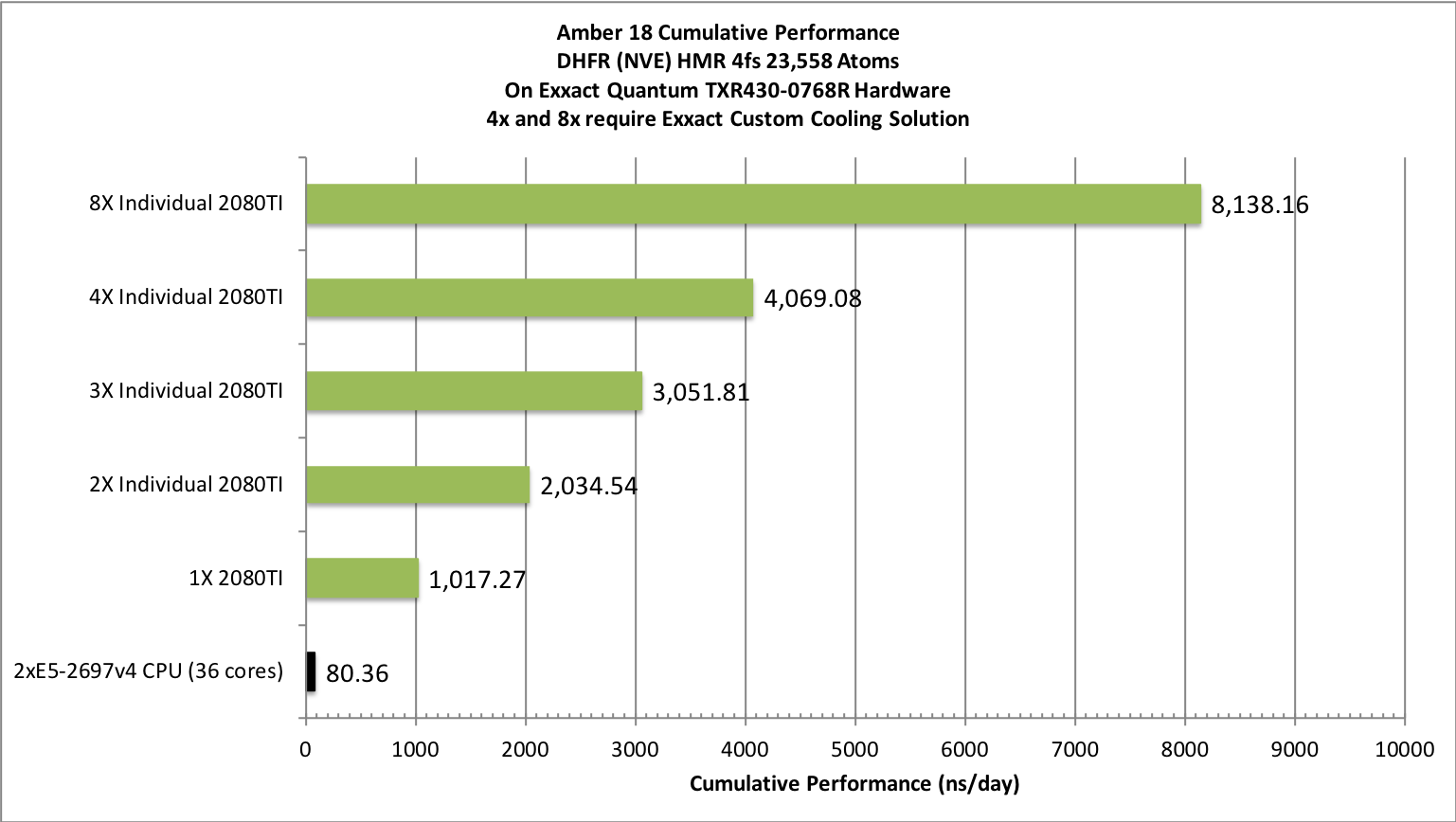
The pmemd.cuda GPU Implementation

Fast and flexible gpu accelerated binding free energy calculations within the amber molecular dynamics package - Mermelstein - 2018 - Journal of Computational Chemistry - Wiley Online Library

Amber (PMEMD) GPU Support
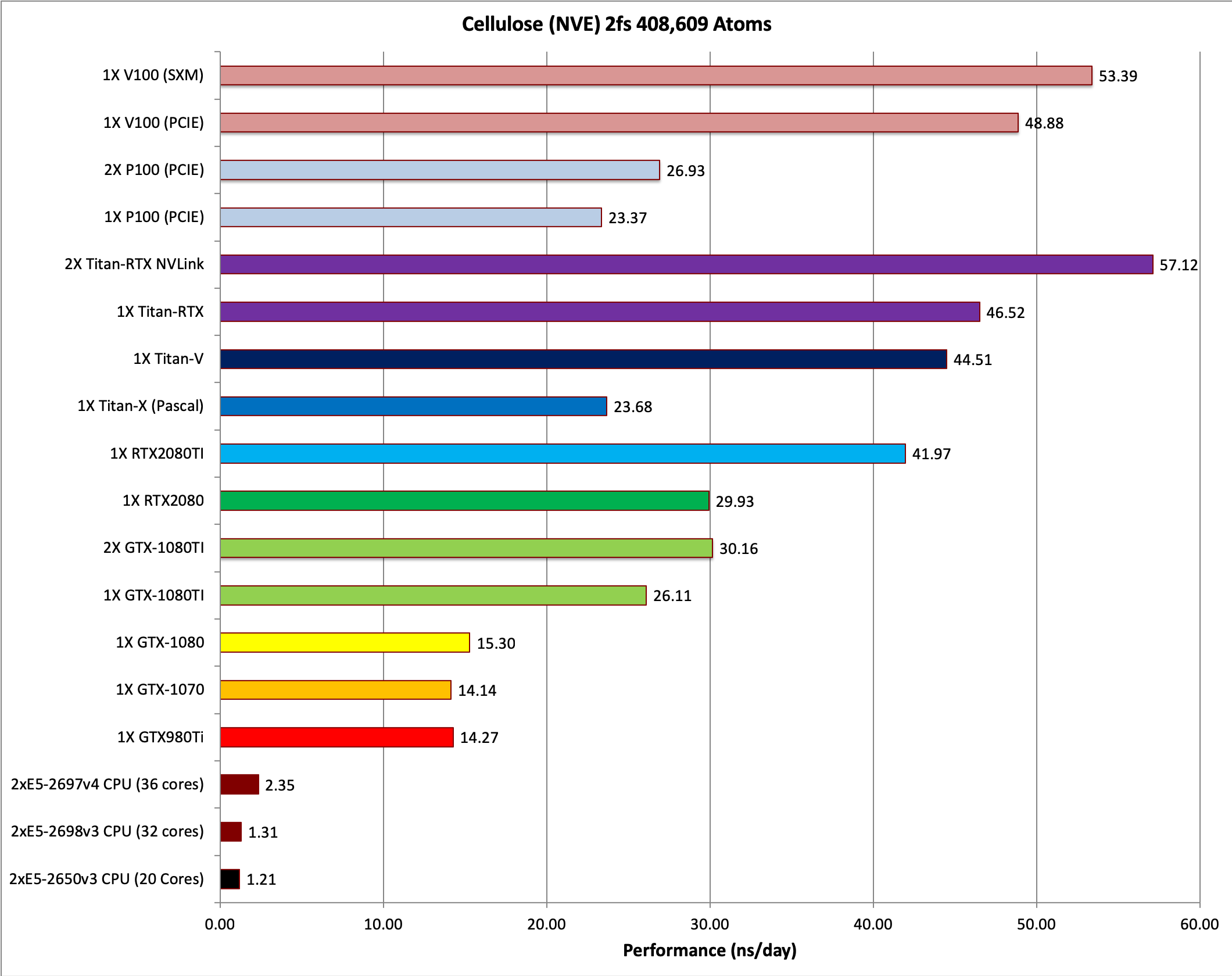
The pmemd.cuda GPU Implementation

Routine Microsecond Molecular Dynamics Simulations with AMBER on GPUs. 2. Explicit Solvent Particle Mesh Ewald
de
por adulto (o preço varia de acordo com o tamanho do grupo)